There’s more than meets the eye when it comes to ‘junk’ DNA

In September 2012, as the final papers from the ENCODE (ENCyclopedia Of DNA Elements) project rolled out, the mood among many molecular biologists was triumphant. After more than a decade of coordinated experiments across dozens of laboratories, the consortium made a bold announcement: a whopping 80% of the human genome showed signs of biochemical activity. Long stretches of non-coding DNA, which had earlier been dismissed as ‘junk’, actually seemed to be humming with life – binding proteins, producing mRNAs, and shaping the behaviour of other genes. “Junk DNA is Dead!” claimed one of dozens of newspaper headlines around the world. Eric Lander, a luminary of the Human Genome Project, hailed ENCODE as the “Google Maps of the human genome.”
But not everyone was celebrating. Dan Graur, an evolutionary biologist at the University of Houston, USA, read the claims with mounting disbelief. To him and many others, ENCODE had made the dangerous mistake of equating molecular “busy”-ness with real biological function and purpose. “We beg to differ,” Graur shot back in a critique in 2013. “ENCODE is considerably worse than even Apple Maps.”
What followed was not a quiet academic disagreement, but an open brawl: barbed blog posts, scathing papers laced with sarcasm, and a debate that spilled well beyond the pages of journals.
At the core of this brawl were two deeply different views of the genome. One side saw extensive biochemical activity as evidence that a large part of our DNA isn’t ‘junk’ as widely believed before, and is extremely functional and finely regulated. The other side, however, envisioned a genome shaped largely by evolutionary accidents, where activity does not automatically mean importance or function.
The brawl also brought into sharp focus a debate that had been unfolding for decades: Can ‘junk’ DNA, that does not code for any protein, even be useful? At the same time, questions intensified after the Human Genome Project confirmed, in 2001, that barely 2% of human DNA encoded proteins. Was 98% doing ‘nothing’?
More than two decades later, scientists in the field are still struggling to pin down the answer to a deceptively simple question: What does it even mean for DNA to ‘do’ something?
Shaped by evolution
The idea of ‘junk’ DNA emerged in the late 1960s from a group of evolutionary biologists who were grappling with a puzzle: Why do some organisms have vastly larger genomes than others – containing far more DNA base pairs – without the organism appearing any more complex?
To repeat a cliché, DNA is the “blueprint” of life. It instructs cells to make proteins that provide structure, catalyse reactions, transport molecules, send signals inside and outside the cell, and defend cells against pathogens. At a higher level, what proteins are made determines what the organism looks like and functions as a whole – its phenotype.
Over time, ‘neutral’ sequences could accumulate, bloating the genome without adding new functions. Ohno famously referred to this excess as ‘junk DNA’
When cells divide, DNA is duplicated and distributed equally between the two daughter cells – but sometimes errors during this duplication can create mutations, which alter the sequence of DNA. If this happens in a sequence that codes for a critical protein, it could hamper the protein’s function and, in effect, the organism’s survival. Sometimes, a mutation could surprisingly make the protein more efficient at its job, boosting the organism’s survival. Thus, random mutations create variations in a population on which natural selection can now act – mutations that are harmful don’t stick around in the genome (because the organism dies), and those that are helpful stay around for a longer time, showing up in more and more organisms as they reproduce.
When scientists were studying ‘simple’ organisms like lungfish and salamanders, they were puzzled to find that these have genomes many times larger than that of humans. If DNA was mainly a set of instructions for building an organism, this made little sense. The complexity of an organism should scale with the length of the instruction manual, yet nature seemed to be handing out the longest manuals to some of its simplest characters. Something else had to be going on.
The onion has feelings? (The C-value paradox)
“The onion test is a simple reality check for anyone who thinks they
can assign a function to every nucleotide in the human genome.
Whatever your proposed functions are, ask yourself this question:
Why does an onion need a genome that is about five times larger
than ours?”
– T Ryan Gregory, Canadian evolutionary biologist
For a long time, biologists naively assumed that the amount of DNA an organism has should roughly track how ‘complex’ it is – more moving parts should mean more DNA. Humans, with our big brains and existential crises, should be right near the top.
Enter the onion. When scientists began measuring genome sizes in the mid-20th century, they discovered that the common onion (Allium cepa) carries about five times more DNA than a human being. Not five times more genes, just more DNA. This discovery became a running joke in genetics, often called the ‘onion test’: if more DNA meant biological sophistication, then onions should be composing symphonies and filing tax returns.
This striking lack of correlation between genome size (called the C-value) and organismal complexity is known as the C-value paradox. It suggests that large portions of DNA could vary in the total amount between species without obvious consequences for complexity, anatomy, or intelligence. Salamanders, lungfish, lilies, and onions all seem to be hoarding DNA, while humans and other mammals make do with relatively streamlined genomes. Whatever all that extra DNA is doing, it clearly isn’t turning onions into philosophers.
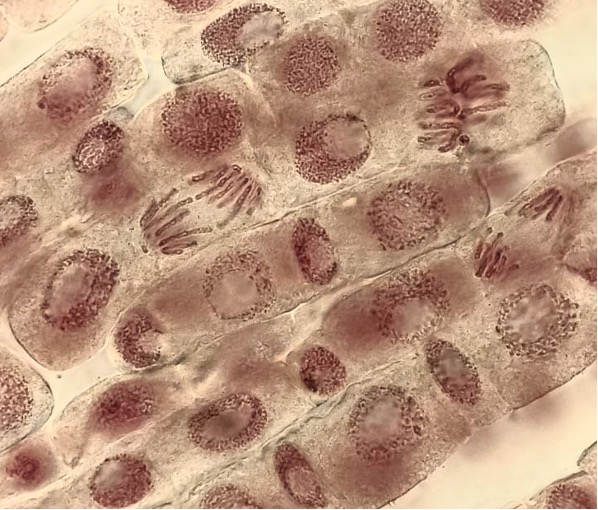
(Photo caption: Microscope image of dividing onion root tip cells, with chromosomes visible as dark threads being pulled apart. Credit: Ryan Ray)
One of the first to articulate this mismatch was Japanese-American geneticist Susumu Ohno. In 1972, Ohno proposed that much of the DNA in large genomes was simply not doing anything useful. His argument was based on evolutionary logic, and the existing observations that organisms have large amounts of repetitive and highly variable DNA. If a DNA sequence does not affect an organism’s survival or reproduction, natural selection has little reason to remove it. Over time, such ‘neutral’ sequences could accumulate, bloating the genome without adding new functions. Ohno famously referred to this excess as ‘junk DNA.’
“The term ‘junk’ suggests a legacy of something that was of use, has no immediate use now, but may acquire a useful function in the future, which is what Ohno meant,” explains Subhash C Lakhotia, Distinguished Professor at Banaras Hindu University. Meanwhile, around 1980, Francis Crick and others published a series of papers in Nature calling this “selfish” DNA. “They thought that although this DNA is of no use to the host cell in which it stays, it can’t be thrown out. And because it’s selfish, it just continues to replicate and stay in the genome,” Subhash adds.
Imagine repeatedly copying a long document by hand, over generations. Glaring errors – a missing paragraph or a garbled sentence – would be corrected immediately. But tiny changes – an extra space or a duplicated word – slip through because they don’t change much in the grand scheme of things.
In the late 1960s, Japanese biologist Motoo Kimura proposed his “neutral theory of molecular evolution,” arguing that much of DNA evolves in exactly this way. Most mutations are neither beneficial nor harmful; they are effectively invisible to natural selection. Genomes accumulate these neutral changes simply because there is no strong pressure to remove them. Duplicate copies of genes that lose function still continue to persist in the genome. Other sequences arrive as genomic hitchhikers inserted by viruses into our DNA. Special sequences called transposable elements copy and paste themselves across chromosomes. The result is a genome which looks less like a blueprint and more like a patchwork quilt, stitched together over time using scraps.
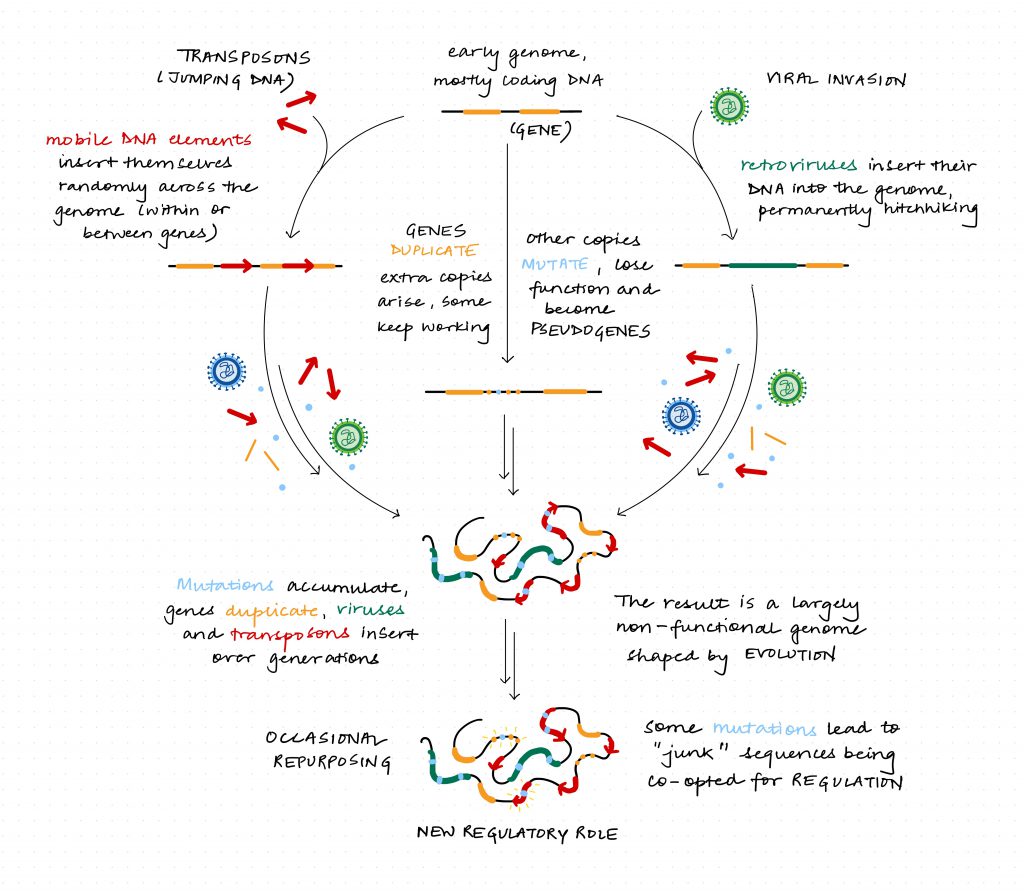
From this perspective, junk DNA is not a mistake. It is exactly what you expect when evolution works without a quality-control engineer. Natural selection is powerful, but it is not perfect – it does not work like an engineer; it works like a tinkerer, as French biologist François Jacob put it.
Evolution, the tinkerer
When biologists talk about the genome as messy or cluttered, they aren’t really accusing nature of incompetence. They are describing how evolution actually works. In his 1977 essay titled Evolution and Tinkering, French biologist François Jacob argued that natural selection does not behave like a careful engineer working from a blueprint. Instead, it improvises with whatever happens to be lying around. “It works like a tinkerer … using pieces of string, fragments of wood, old cardboard – odds and ends – to produce something that merely works,” he wrote.
This tinkering leaves behind a genome full of redundancies and long stretches of DNA that appear unnecessary. The result is not elegance or efficiency, but adequacy. Once something works well enough to pass on genes, there is little pressure to tidy up the rest – and the genome is inherited, patched, and passed on, piling up clutter over generations.
“Evolution is shaped by randomness and unpredictability,” Moran wrote in his book What’s in Your Genome? “Genomes reflect that – they are sloppy and not exquisitely designed like a Swiss watch.”
In organisms with low population sizes, where there is little competition for space and resources, even slightly deleterious mutations can be just as easily maintained over generations as neutral ones, increasing the amount of DNA that doesn’t ‘do’ anything. “Most species with large genomes today have had low population sizes for much of their history, and that explains why they can have lots of slightly detrimental junk DNA,” says Laurence Moran, Professor Emeritus at the University of Toronto.
For decades, this view was widely accepted because there was no way to systematically test what non-coding DNA was doing. The term ‘junk’ represented non-functional, not useless. But soon, that changed when new technologies started revealing something unusual across the genome.
Identical genomes, distinct cells
Around the same time as Ohno and Kimura, scientists started realising that genes were only a small part of what actually shapes a cell. After all, if all the cells of our body carry the same genes, how could they behave so differently across tissues, developmental stages, and environments?
The first semblance of an answer began to appear in the 1960s and 1970s, when geneticists began discovering that genes were surrounded by mysterious control regions. In 1961, François Jacob and Jacques Monod proposed the idea of gene regulation – certain DNA sequences outside the genome could turn genes on and off, controlling whether and when they made proteins. It was an early hint that some DNA elements could have roles beyond coding for proteins.
In the 1980s and 1990s, scientists began uncovering DNA elements scattered across the genome that act like switches. Enhancers, first identified in the 1980s, were often found to sit far from the genes they control, sometimes hundreds of thousands of base pairs away. These sequences loop through space to bind to genes and boost their activity in specific tissues. A single enhancer can control whether a gene is active in the brain but silent in the liver. Suddenly, vast stretches of non-coding DNA were revealed as a regulatory wiring system.
Parallely came the RNA revolution. For a long time, the only thing we knew (and believed) was that DNA is converted into mRNA, an intermediate messenger molecule, by a process called transcription in the nucleus of cells. mRNA then moves out of the nucleus and is translated to make proteins.
But in the 1960s, scientists started finding RNA that stayed in the nucleus and never exited into the cytoplasm. “Since translation can only happen in the cytoplasm, the implication was that these RNAs don’t get translated,” explains Subhash. “But then the bigger question was: What are they even doing there?”
By the early 2000s, hundreds of non-coding RNAs were discovered in humans, plants, and animals, acting as molecular dimmer switches that fine-tune gene expression
The question remained unanswered until 1993, when American scientists Victor Ambros and Gary Ruvkun studying tiny worms made the startling discovery that small RNA molecules – later called microRNAs – could control gene activity without ever becoming proteins. What had once been dismissed as transcriptional noise turned out to be a powerful regulatory system. By the early 2000s, hundreds of these non-coding RNAs were discovered in humans, plants, and animals, acting as molecular dimmer switches that fine-tune gene expression.
Non-coding DNA and RNA also shape the physical structure of the genome inside the cell. DNA isn’t laid out like a straight line – it folds, loops, and packs into three-dimensional forms. “Some non-coding sequences act as anchors and scaffolds for this folding, helping organise the genome in space so that the right genes can interact with the right regulatory elements,” explains Saumitra Das, Professor at the Department of Microbiology and Cell Biology, IISc. Other sequences can influence how tightly a DNA sequence is folded up.
Don’t shoot the gene, blame the switch
In the late 1990s, geneticists were puzzled by families in which children were born with extra fingers or toes – a condition known as polydactyly. Genetic tests came back clean. The SHH gene, which plays a crucial role in limb development, was completely normal. There was no broken protein, and no obvious culprit.
The actual culprit turned out to lie nearly one million DNA base pairs away from the SHH gene – in a stretch of non-coding DNA. This was the ZRS enhancer sequence, which acts like a remote switch and turns the SHH gene on at the right place and right time during embryonic development. In patients with polydactyly, tiny mutations in the ZRS enhancer caused SHH to switch on in the wrong place. The gene itself worked perfectly, but it was expressed where it shouldn’t have been. The result was an extra signal telling the limb to grow additional digits.
This added weight to the growing recognition that a disease isn’t always caused by a mutated gene making a broken protein. “In many cases, the protein itself is perfectly intact,” explains Saumitra Das. “What goes wrong is when, where, or how much of it is made.” In other words, the problem often lies not in the parts, but in the switches that control them – and this can have dramatic effects on the phenotype.
Similar regulatory mishaps are now being uncovered across medicine. In certain cancers, subtle changes in regulatory regions or non-coding RNAs can push growth genes into overdrive. In neurological and autoimmune disorders, genes may be activated too early, too late, or in the wrong cells altogether. “For patients and clinicians, this has expanded the diagnostic searchlight beyond genes alone,” says Saumitra. This is also a stark reminder that junk DNA may or may not matter much, depending on which side of the debate you are – but when it matters, it can matter profoundly.
These functions of non-coding DNA may seem intuitive today, but for a long time, they were anything but obvious, which made studying them harder.
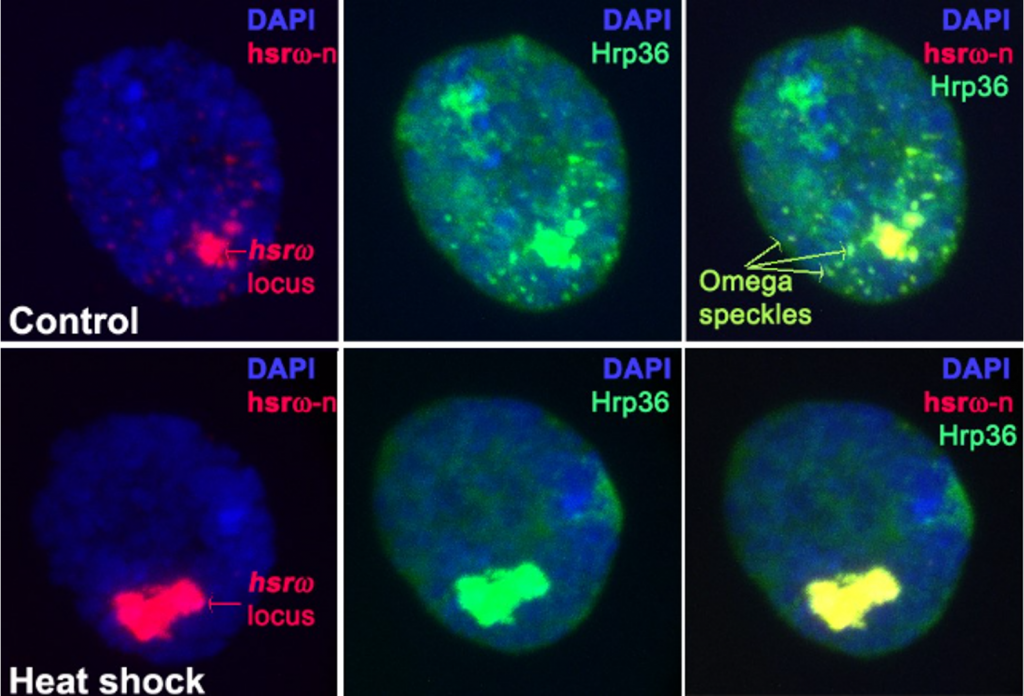
Subhash underscores this with an example. Much of his earlier work was on a gene in the fruit fly Drosophila called hsrω. “I was looking for what protein it makes, and then it turned out it doesn’t make a protein at all,” he narrates. This was in the 1980s, around the same time that Crick first talked about selfish DNA, and hsrω seemed like it fit the brief. “But I didn’t believe that this was a selfish gene, and I kept on pursuing it – and people kept asking me why,” says Subhash. “My competitors abroad, who had started working on this gene because of our initial work, had to give up because they wouldn’t get funding to work on this seemingly selfish gene. But fortunately, I did.”
What Subhash found was that hsrω makes multiple long non-coding RNAs which could bind to a variety of RNA-binding proteins called hnRNPs, making these proteins unavailable for binding to their real targets and regulating whether or not they would be available to perform their functions. Over the next few decades, Subhash uncovered other functions of hsrω in development and stress responses.
But even though evolutionary biologists have never really denied that some non-coding DNA matters, they were still wary of attributing “usefulness” to it. And when the Human Genome Project announced its findings, things really came to a head.
Hunting for purpose
At the turn of the millennium, something changed dramatically. When the first draft of the human genome sequence was published in 2001 with a lot of fanfare, it was the first time scientists could see the genome as a whole. It was expected to settle long-standing questions about genetic complexity. Many anticipated millions of genes to exist. Shockingly, the number turned out to be closer to a mere 20,000 – not much more than in the genome of a worm or a fly.
The surprises didn’t end there. Only about 2% of the genome, it seemed, coded for proteins. The remaining 98% was composed of repeated sequences, duplicated fragments, and vast stretches with no obvious role. Far from undermining the idea of junk DNA, the genome sequence seemed to confirm evolutionary predictions – large, complex organisms carried large amounts of DNA that did not appear to be under the influence of natural selection.
After the Human Genome Project, evolutionary biologists continued to be cautious, but molecular biologists began to push a bold narrative … they claimed that perhaps very little of the genome is truly junk at all
This abundance of ostensibly useless DNA created both opportunity and unease. Evolutionary biologists continued to be cautious, but molecular biologists began to push a bold narrative: as evidence mounted for the importance of non-coding DNA, they claimed that perhaps very little of the genome is truly junk at all.
Individual examples of non-coding function were already well established, but they were discovered piecemeal, one system at a time. What was missing was a genome-wide view. As new technologies made it possible to survey the entire genome at once, a new ambition to move beyond sequence, map activity, and probe regulatory and structural roles of non-coding DNA emerged.
From this excitement, ENCODE was born. Launched in 2003 by the US National Human Genome Research Institute, the project brought together hundreds of researchers across dozens of labs. It aimed to catalogue every detectable functional element in the human genome using a uniform experimental framework. ENCODE set out to annotate the DNA sequence that had come out of the Human Genome Project – mapping where proteins bind, where RNA is produced, and which regions act as regulatory switches. What they sought was clear proof of usefulness or “function” for non-coding DNA.
But the problem was that ENCODE’s definition of function was somewhat vague. According to the researchers, for a DNA segment to be “functional,” it needs to be either transcribed by RNA, bound by transcription factors (the proteins that control gene expression) or be marked by regulatory signals and modifications (like methylation). If a DNA segment binds to transcription factors, it is functional. So, other DNA segments that bind to transcription factors should also be functional. This is what they presumed.
Driven by these ideas, and after a decade of large-scale experiments, the ENCODE consortium published a coordinated set of more than 30 peer-reviewed research papers in Nature, Science, and Genome Research in 2012. Together, these studies mapped patterns of transcription, protein binding, chromatin modification, and DNA accessibility across dozens of human cell types.
Based on the prevalence of this biochemical activity, the consortium concluded that 80.4% of the human genome showed evidence of “function.” In an accompanying editorial, Stella Hurtley, a senior editor of the journal Science wrote, “No More Junk DNA.”
The function wars
At the University of Houston, Dan Graur was upset. ENCODE’s statement went against decades of evolutionary theory. According to the junk DNA model, it was the other way around: Only about 10% of the genome is evolutionarily conserved across species and selected for – which should be the functional portion – with the remaining 90% being ‘junk’. “ENCODE’s estimate of 80% [functional DNA] implied that at least 70% of the genome is invulnerable to harmful mutations,” Graur wrote in a journal article in Genome Biology and Evolution in 2013. “Either because no mutation can ever occur in these so-called functional regions, or because no mutation in these regions can ever be deleterious.”
Graur famously compared this to claiming that a television set left on and unattended will still be in working condition after a million years because no natural events, such as rust, erosion, static electricity, and earthquakes, can affect it.
Just because a sequence does something does not prove a biological purpose, and it certainly does not mean that the organism would suffer if it were removed
This is absurd – clearly, there was a flaw in ENCODE’s argument. A DNA segment can display a property without necessarily having the associated function. A random sequence may bind a transcription factor, but that may not result in transcription into RNA. Cells are noisy places; enzymes bind DNA promiscuously, RNA is transcribed and degraded without consequence. Just because a sequence does something does not prove a biological purpose, and it certainly does not mean that the organism would suffer if it were removed.
Thus began the function wars, marked by blunt critiques, sharply worded papers, and an unbridled willingness among scientists to be publicly, and unapologetically, combative.
“In the absence of any solid evidence that a given sequence is functional, the default explanation has to be that it is not,” believes Moran. This is why countless philosophers and experts in molecular evolution were upset with the 2012 ENCODE papers, and it is why they were retracted in 2014. Unfortunately, most scientists didn’t read this retraction.
“A large part of the problem isn’t about biochemical activity. It’s about a misunderstanding of what the debate is about,” explains Moran. “Dozens of papers that talk about non-coding DNA today say that scientists like Susumu Ohno and Motoo Kimura were really, really stupid.”
According to this modern caricature, Ohno and Kimura – writing in the 1960s and 1970s – supposedly believed that the only functional DNA in the genome was protein-coding DNA, and therefore dismissed everything else as junk. Contemporary authors then show that a tiny bit of non-coding DNA has a function and claim that this disproves the entire concept of junk DNA.
“The ignorance of these modern scientists is frustrating,” Moran says vehemently. “The original proponents of junk DNA were not stupid. They knew about all kinds of non-coding DNA that is functional. No knowledgeable scientist ever said that all non-coding DNA was junk, and anyone who makes such a false claim is simply demonstrating their ignorance of history.”
When the ENCODE results were published in 2012, John Stamatoyannopoulos, a leading figure in the consortium, believed that their findings would necessitate the rewriting of textbooks. “We agree,” Graur retorted in 2013. “Many textbooks dealing with marketing, mass-media hype, and public relations may well have to be rewritten.”
All of this matters because language shapes expectations. To the public, and even to young researchers, the idea that almost nothing in the genome is waste can smuggle in a comforting but misleading assumption: that biology is optimised, tidy, and intentional. Evolutionary biologists have long argued the opposite. The genome is not a masterpiece engineered from scratch, and is, in fact, full of redundancies and leftovers. When this messiness is flattened into a story of near perfection, something essential about how evolution works is lost.
“The general public and many scientists believe that humans are designed. But people should know that there’s another worldview out there – composed of people like me who do not think the world looks designed,” says Moran. “We think the world looks sloppy and inefficient, and that this is exactly what you would expect if life evolved.”
Parth Kumar is a third year BSc (Research) student at IISc and a former science writing intern at the Office of Communications
(Edited by Ranjini Raghunath)