Cryo-EM, an emerging technique, is allowing scientists to better zoom in on cells and molecules
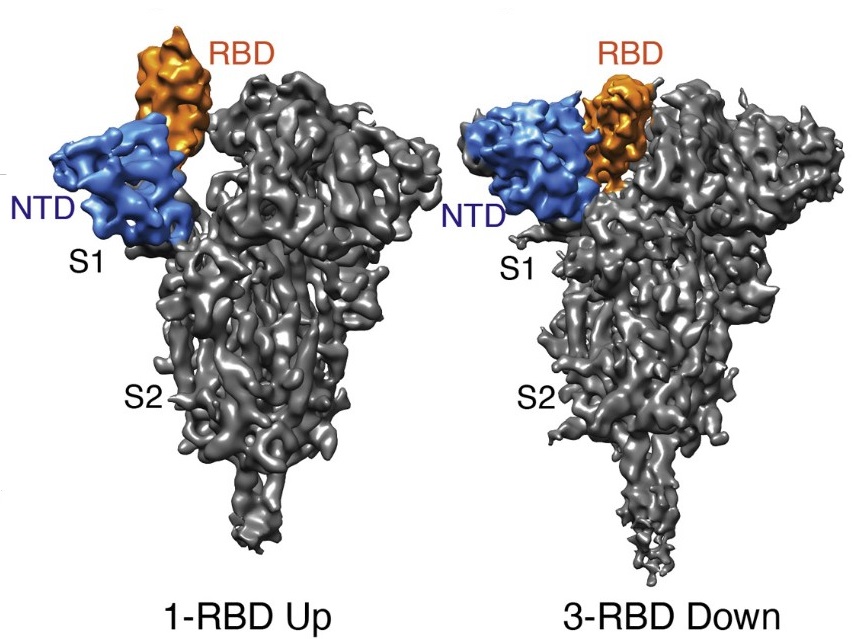
“Like any living organisms, we are a bag of water, formed from billions of cells which all are small bags of water. Since air is not transparent to electrons, an electron microscope must operate under vacuum – which means that any observed biological specimen must be dry.” This opening line of the Nobel lecture given by Swiss biophysicist Jacques Dubochet in 2017 perfectly sums up the problem in using electron microscopy to observe biological specimens. Most of us are familiar with the upright microscopes we used in schools to peer inside a tissue sample mounted on a glass slide. These microscopes need a source of light – which is either sunlight or a lamp. An electron microscope operates on fundamentally similar principles to those of light microscopes, except for one key difference – instead of visible light, electron beams are used to visualise the object. The reason for using complicated electron beams where simple light could have done the trick is resolution – the ability to distinguish between two closely situated points. One nanometer (nm) is one-billionth of a metre, and the resolution of a light microscope is about 200 nm. This is good enough to distinguish large organelles (compartments) inside the cells. But an electron microscope has a far higher resolution. At 0.1 nm, it is good enough to distinguish between individual atoms.
When we think of a microscope, what comes to mind is looking at a section of a leaf or an animal tissue under its lenses. The use of a microscope is fundamental to the study of biology, and naturally, having the resolving power of an electron microscope would provide insights at an atomic level. But it took a very long time for an electron microscope to be commonly used to visualise biological specimens. Any part of a living organism is at least 70% water. And as Dubochet says, electron microscopes need to be operated in vacuum, devoid of any moisture. This makes biological specimens fundamentally incompatible for studying under an electron microscope – locking away the secrets of life at an atomic and molecular level from the scientists. This is the reason that although the electron microscope was developed in the 1930s, for a long time it was only used to look at atomic details of inanimate materials, like various metal alloys.
It wasn’t until over half a century later, in the 1990s, that scientists around the globe began to use it routinely to look at molecules – mostly proteins – that shape our life. This change came about through the development of a revolutionary new technique of sample preparation which led to the term – “cryogenic electron microscopy” or Cryo-EM for short, for which Dubochet, alongside Joachim Frank and Richard Henderson, won the Nobel Prize in chemistry in 2017.
Bringing electron microscopy to biology
Visualising biological specimens with an electron microscope came with a myriad of problems. First, as mentioned earlier, an electron microscope must be operated in vacuum and these specimens don’t hold up very well in vacuum due to their high water content. Secondly, biological samples are fragile. The high energy electrons would collide with and potentially destroy them – and hence they can only be observed under a very low energy electron beam. This significantly affects the quality of the image obtained. It took our three laureates years of work to overcome these challenges.
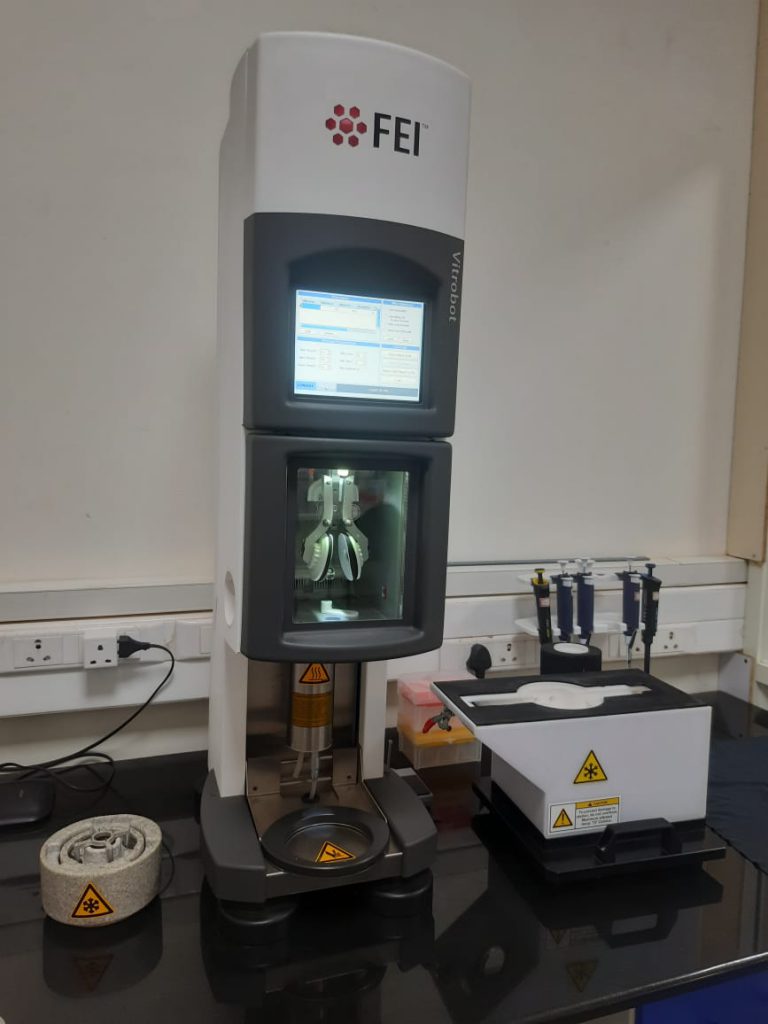

Dubochet was the person responsible for the “cryo” in Cryo-EM. He was the one who developed a special method of sample preparation that allowed biological samples to be examined by an electron microscope. One workaround to the problem of having moisture-rich samples would be to freeze them. But water, upon freezing, forms ice crystals, where the molecules are well structured in a cage-like fashion. Such ordered ice crystals form a barrier to the passage of electron beams. Dubochet used liquid nitrogen to flash-freeze biological samples suspended in liquid ethane. Doing this freezes the samples so fast that water molecules do not have time to form highly ordered ice crystals, and instead, they form a disordered glass-like structure, which allows the electron beams to easily pass through. Flash-freezing also solves the problem of having any free moisture in vacuum. It also preserves the biomolecules in near-original condition, allowing scientists to look at biological processes frozen in time.
As mentioned earlier, biological samples have to be subjected to the lowest energy electron beam to ensure sample integrity. This means that plenty of processing post-microscopy is required to obtain a sharp, high-resolution image. Enter Joachim Frank, whose image processing algorithms made it possible to compile 2D pictures of a protein taken using Cryo-EM and build a sharp 3D structure with details at atomic levels.
Richard Henderson was the first person to successfully generate an image of a protein, a bacterial pigment called rhodopsin, using an electron microscope, and he kept refining his technique till he obtained a protein structure with the same resolution as that obtained using another popular method – X-ray crystallography.
Catching proteins in action
Since the development of Cryo-EM techniques, their most widespread use has been to look at proteins and characterise their structures. But structural biologists already had other tools like X-ray crystallography and Nuclear Magnetic Resonance (NMR) at their disposal to do exactly that. So, what was special about this new technique? First, both X-ray crystallography and NMR require huge quantities of the protein whose structure is to be determined, which can be a bottleneck in research, while Cryo-EM requires a very small amount. Secondly, Cryo-EM allows scientists to capture snapshots of protein structures while they are carrying out biological processes inside cells, which is not possible with either of the other techniques, as they require proteins to be purified from the rest of the cell. Having Cryo-EM as a tool at their disposal, it has become possible for scientists to understand how protein structure changes as it interacts with drugs, pathogens, and other molecules inside the cell.
“While X-ray crystallography and NMR still remain popular techniques, Cryo-EM is rapidly becoming more and more commonly used”
While X-ray crystallography and NMR still remain popular techniques, Cryo-EM is rapidly becoming more and more commonly used, thanks to the development of better image processing software, more stable microscopes, improved back-end support, improved cameras to capture images and so on. In 2020, the 10,000th protein structure determined by Cryo-EM was deposited in a prominent database for protein structures.
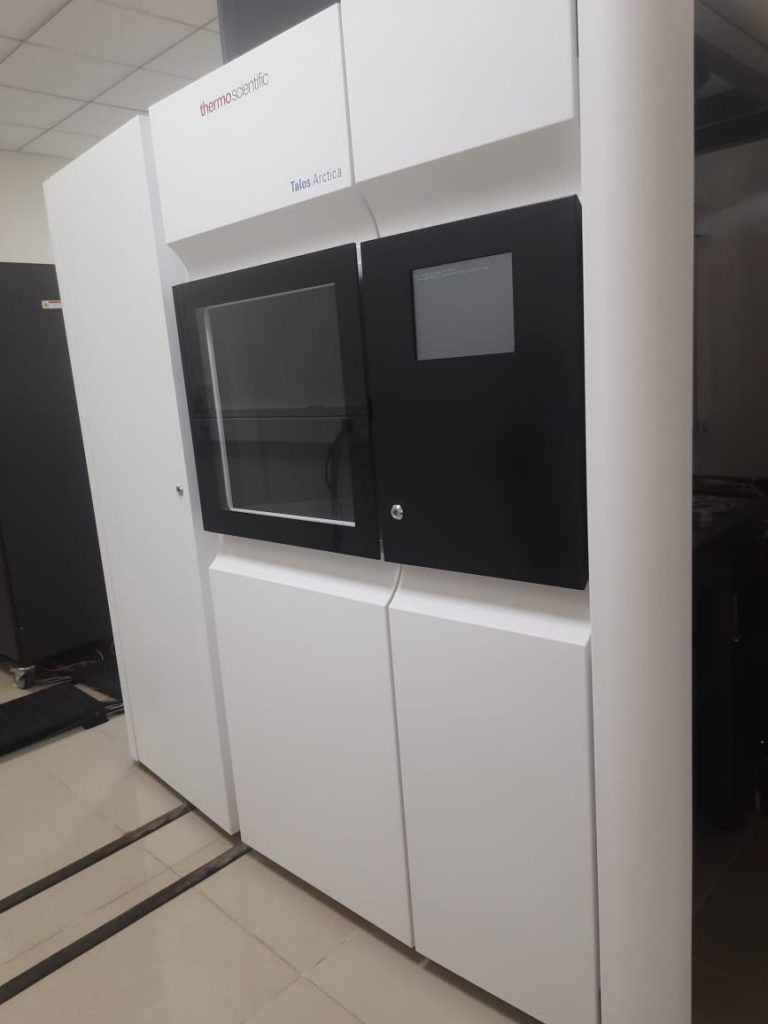

Cryo-EM has become particularly popular for determining structures of membrane proteins. Each of our cells is bound by a double-layered membrane made of molecules called lipids. This lipid membrane is often interrupted by proteins embedded in it. Proteins in the membrane may form channels that allow molecules from outside to enter the cell, or receptors that interact with “signalling” molecules that provide the cell with information about the external environment. Such membrane proteins are particularly difficult to purify in large quantities as they prefer living in the water-free environment of the lipid bilayer and are not easily soluble in water. Cryo-EM is a good tool not only to decipher the structures of these proteins, but also to understand how their structures change, and how the proteins interact with other molecules. Some of the molecular movements and interactions that have been understood using Cryo-EM include the transport of water or ions inside cells, and the effects of drugs and pathogens on a cell. Being able to answer such questions provides crucial insights into how biological systems work.
Cryo-EM in India
It was 2017 when Dubochet, Frank and Henderson were recognised for their accomplishments with a Nobel Prize. The following year, in 2018, two Cryo-EM facilities opened their doors right here in Bangalore – one housed at IISc and the other at the National Centre for Biological Sciences (NCBS). The primary goal of these facilities was to expose the faculty and student researchers of their respective host institutes to the latest frontiers in Cryo-EM. The facilities also provide training and guidance, and allow academic and industrial researchers from other institutes to use them as well.
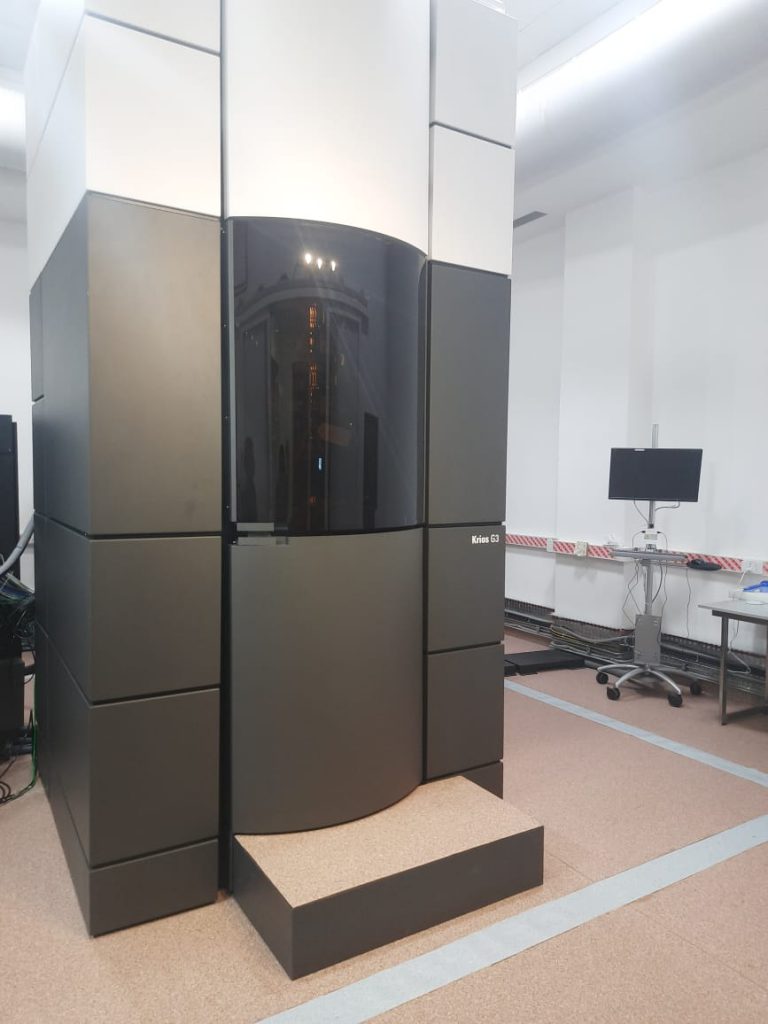

The Cryo-EM facility at IISc was established with efforts led by Somnath Dutta, Associate Professor at the Molecular Biophysics Unit with the help of Raghavan Varadarajan, the then MBU Chair. Somnath’s journey in cryo electron microscopy has been long, adventurous, and enjoyable, he says – starting from his PhD days at the National Institute of Cholera and Enteric Diseases (NICED), Kolkata. He studied under the supervision of retired faculty member AN Ghosh who pioneered the use of this technique in India, back in the 1990s, with the help of one of the first cryo-electron microscopes in the country. For his PhD, Somnath was studying toxins from cholera bacteria when he saw a structure like a “bead necklace”, under the electron microscope. That was the beginning of his lifelong fascination with electron microscopy. “I enjoy imaging and photography a lot, and Cryo-EM allows you to take images at a microscopic level,” says Somnath. When he joined the Institute in 2016, he was interested in looking at drugs that interact with bacterial toxins in the biological membrane. Cryo-EM would have been an extremely useful technique for this purpose, however, at that time IISc did not have this facility. Due to his persistent efforts, he explains, the Advanced Center for Cryo Electron Microscopy became operational two and a half years down the road.
From only a handful of labs using this technique in the 1990s and early 2000s, to having fully operational world-class facilities in multiple institutions today, Cryo-EM has come a long way in India
These days, multiple labs, both within and outside of IISc, use this facility to elucidate structures of proteins, or to peer into cells at an atomic resolution. It is also used by researchers in the pharma industry who study drug interactions at molecular levels to gain approval for commercialising their product from regulatory bodies like the Central Drugs Standard Control Organisation (CDSCO). As for Somnath himself, his lab is mostly interested in studying secretion systems of bacteria, bacterial toxins and other membrane proteins. In some of the recent studies published by his group, they have determined, visualised, and characterised the structure and function of biological macromolecules involved in host-pathogen interactions, which are associated with infectious diseases including SARS-CoV-2 and tuberculosis, using Cryo-EM techniques.
Around the same time that the facility was planned to be set up at IISc, Vinothkumar Kutti Ragunath, faculty member at NCBS, was working with his colleagues to set up a similar facility at his institute in 2017-2018. Like the facility at IISc, this one too is open to researchers across the country. Students using the facility also get the opportunity to obtain hands-on training in using the electron microscopes. “Students should be confident in using the instrument beyond merely collecting data,” believes Vinoth. His own journey with Cryo-EM also started during his PhD days at the Max-Planck Institute of Biophysics, Frankfurt. At that time, he too was primarily interested in looking at membrane proteins. He recalls that among the techniques available at that time, crystallisation of membrane proteins was a difficult endeavour. The proteins would often go into inclusion bodies (clumps) or express very little, and it was often difficult to purify in large quantities. One of the many advantages of single particle cryo-electron microscopy is that it does not require crystallisation and is easier to perform with lesser quantities of protein. His group currently studies various membrane proteins and enzymes using Cryo-EM.
From only a handful of labs using this technique in the 1990s and early 2000s, to having fully operational world-class facilities in multiple institutions today, Cryo-EM has come a long way in India. Vinoth attributes this rise in popularity to a “series of developments happening in the background starting from early 2000s to 2011-12,” most prominent of which is the development of better detectors and algorithms that can capture and analyse the electrons when they hit the sample.
Beyond proteins
Cryo-EM is not used solely to determine protein structures, it is also being used by many scientists to look at samples of intact cells – a process called cyto-tomography. As Somnath remarks, “electron microscopy is first and foremost an imaging technique,” and it should be used as one – to go beyond determining atomic structures and start investigating the bigger picture.
Cyto-tomography has been used to look at tiny viruses infecting cells, at biological processes, like cells internalising water and ions through channels on the membrane, or how drug molecules interact with organelles inside cells at single molecule resolution, and more. It can be used to visualise how different molecules interact with each other in cells, what kinds of changes cells undergo when they are dividing or are under attack by a pathogen, and what different structures look like inside the cell. It can also be used in tandem with light microscopy, where staining with light microscopy would give us the exact location of a molecule or organelle, and looking at the same position in cells using cryo electron microscopy would give us a more detailed understanding of the processes going on in that location.
Despite all its advantages, Cryo-EM suffers from some serious limitations. First, since samples have to be frozen, it is impossible to image live cells. Second, there is a lower limit to the size of protein that can be determined by Cryo-EM – anything less than the size of 60 kilodalton has a low signal-to-noise ratio, and hence a clear image cannot be obtained. But with the improvement of detectors and better computational programs, it may become possible in the future to obtain structures of smaller proteins. For now, a clever workaround to this problem is to tack on a larger molecule to the small protein of interest, and then obtain the structure of the whole complex. The structure of the smaller protein can then be deduced from that information.
“Thirty years ago, my group presented a model of a virus floating in immobilised water, showing details as small as 35 Å. Nowadays, cryo-electron microscopists are nearly routinely achieving 3.5 Å. Thirty years later, a resolution 10 times better, a volume 1,000 times smaller; this is truly an impressive achievement,” Dubochet had remarked in his Nobel lecture. He ended by saying, “At 3.5 Å resolution, atoms are visible. We see chemistry, how the atoms are arranged in the molecules, how the disease changes the arrangement. Perhaps we will find which drug disentangles the aggregates that make a brain senile … Colleagues, cryo-electron microscopists, you have a good tool at your disposal; make the best of it!”
Prarthana Ghosh Dastidar is a PhD student in the Department of Microbiology and Cell Biology, IISc and a science writing intern at the Office of Communications